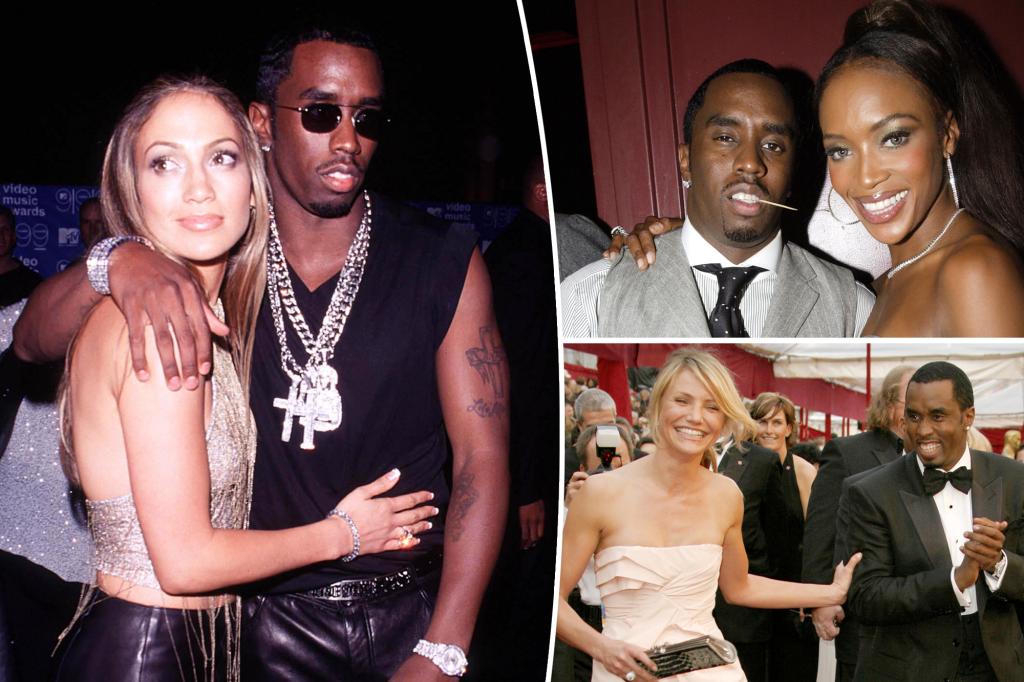
Sean “Diddy” Combs has had a star-studded dating history, being romantically linked to various A-listers over the years, including Jennifer Lopez and Cameron Diaz. He has also been in long-term relationships with Cassie and Kim Porter. Diddy is a father of seven and has maintained close relationships with many of the mothers of his children.
Cassie filed a lawsuit against Diddy in November 2023, which included accusations of rape, physical abuse, and more. After quickly reaching a settlement, Diddy faced multiple sexual assault claims and had his homes raided by Homeland Security in connection to a sex-trafficking investigation. He denied any wrongdoing through his lawyer. Diddy’s high-profile romances have often been in the spotlight due to his fame and the fame of his partners.
Diddy’s dating history includes well-known names such as Misa Hylton, who he dated early in his career and welcomed their first child with. He also had a long-term relationship with Kim Porter, with whom he had several children. Kim Porter passed away in 2018, and Diddy expressed his immense grief over losing what he described as a relationship that was more than just romantic. Diddy has also dated Jennifer Lopez, Naomi Campbell, Sarah Chapman, and Cameron Diaz among others.
Diddy’s relationship with Cassie was tumultuous, beginning in 2007 and ultimately ending in a lawsuit from Cassie in 2023. The lawsuit claimed that Diddy exerted control over her life, leading to lasting trauma. Cassie has moved on and is now married to Alex Fine. Diddy was also briefly linked to Cameron Diaz, Lori Harvey, Yung Miami, Gina Huynh, and Dana Tran. These relationships were often the subject of media attention due to Diddy’s fame and the connections to other well-known figures.
Despite his many high-profile romances, Diddy has continued to face legal challenges and controversies. The rapper has denied any accusations of wrongdoing against him and has sought to move forward with his career and personal life. His relationships have been marked by highs and lows, and he has faced public scrutiny over his actions in some cases. However, Diddy has also expressed love and respect for many of his former partners, highlighting the complexities of his personal life.
Throughout his dating history, Diddy has navigated relationships with some of the most famous women in the entertainment industry. His romances have often been the subject of speculation and media attention, with fans and the public eagerly following his love life. Despite the challenges and controversies, Diddy has maintained his status as a prominent figure in the music industry and has continued to pursue his passions and interests. His dating history reflects the ups and downs of his personal life and the complexities of relationships in the public eye.