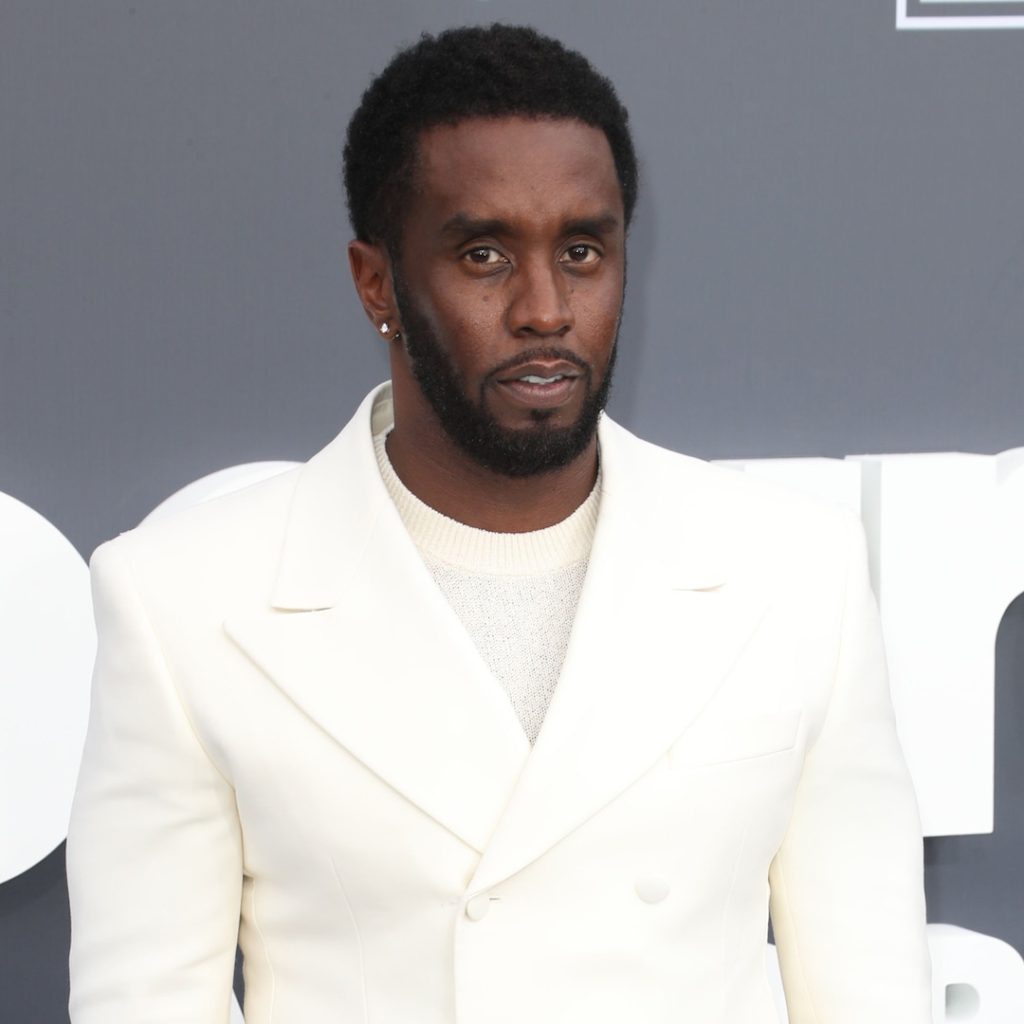
Mr. Combs’ lawyer, Dyer, criticized the use of excessive force during the execution of search warrants at the rapper’s residences, calling it a gross overuse of military-level force. Dyer expressed concern over the treatment of Combs’ children and employees, noting that there was no excuse for the hostility exhibited by authorities. He described the raids as an “unprecedented ambush” but clarified that Combs was not detained during the operation. Dyer also emphasized that Combs had cooperated with authorities and denied any criminal or civil liability.
Dyer labeled the raids as a “witch hunt” and asserted that there had been no findings of criminal or civil liability related to the allegations against Combs. He maintained Combs’ innocence and stated that the rapper would continue to fight to clear his name. Despite the seriousness of the allegations, Dyer seemed confident in his client’s innocence and willingness to cooperate with authorities. E! News reached out to the Department of Homeland Security for comment on Combs’ allegations, but there has been no response from the agency thus far.
The criticism of the use of excessive force and the description of the raids as an “unprecedented ambush” suggest that the authorities’ actions may have been perceived as aggressive and disproportionate. Dyer’s defense of Combs’ innocence and cooperation with authorities indicates a strong belief in the rapper’s character and willingness to address the allegations against him. The lack of response from the Department of Homeland Security leaves unanswered questions about the justification for the methods used during the raids and the nature of the allegations against Combs.
Dyer’s statement raises concerns about the treatment of Combs’ family and employees during the raids, highlighting the impact of the authorities’ actions beyond the rapper himself. The emphasis on Combs’ innocence and commitment to clear his name suggests a determination to address the allegations and defend his reputation. The characterization of the raids as a “witch hunt” implies skepticism about the motives behind the operation and the validity of the allegations against Combs. Overall, Dyer’s statement conveys a strong defense of his client and challenges the conduct of the authorities involved in the investigation.
The lack of response from the Department of Homeland Security to inquiries about Combs’ allegations leaves the situation unresolved and leaves room for further developments in the case. The controversy surrounding the raids and the allegations against Combs may continue to unfold as more information becomes available. Despite the challenges faced by Combs in relation to the raids, his legal representation appears determined to vigorously defend his innocence and reputation. The outcome of this situation remains uncertain, but Dyer’s statement sets the tone for a potentially contentious legal battle ahead.