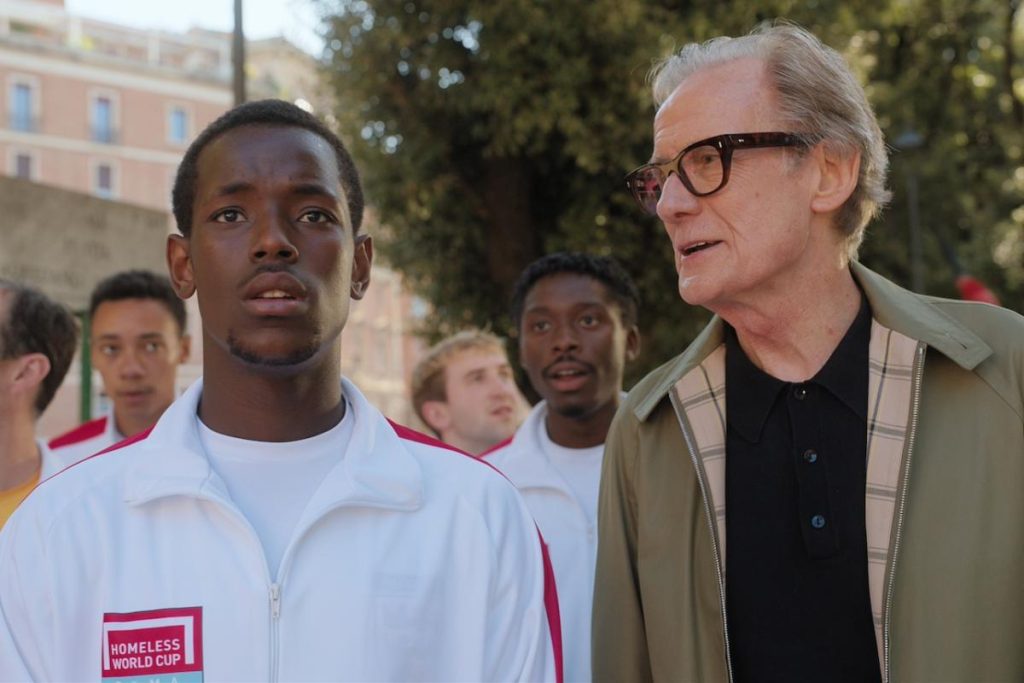
The Beautiful Game, a new sports drama starring Bill Nighy, is now available for streaming on Netflix. Directed by Thea Sharrock and written by Frank Cottrell-Boyce, the film follows the story of a skilled football player named Vinny, played by Michael Ward, who joins a team for homeless players that competes in the Homeless World Cup. While The Beautiful Game is based on the true story of the Homeless World Cup, the characters in the film are fictional.
The Homeless World Cup is an annual international soccer tournament for players experiencing homelessness, organized by the non-profit Homeless World Cup Foundation. Players participating in the tournament must meet specific criteria, such as having been homeless in the past year, being a street paper vendor, or being in addiction rehabilitation. The tournament aims to provide opportunities for individuals facing homelessness to come together and showcase their skills on the field.
The filmmakers of The Beautiful Game worked closely with the Homeless World Cup Foundation to ensure the authenticity of the tournament portrayed in the movie. Director Thea Sharrock consulted with former Homeless World Cup players and ambassadors to accurately depict the event and the experiences of the players. Many of the background actors in the film were real-life former participants, adding to the authenticity of the film.
The character Gabriella, played by Valeria Golino in the film, is inspired by Mel Young, the co-founder and president of The Homeless World Cup. While the protagonist Vinny and his coach/manager, Mal, are fictional characters, their story is meant to showcase the challenges and triumphs faced by individuals experiencing homelessness. The movie captures the spirit of the Homeless World Cup and the impact it has on the lives of its participants.
The Homeless World Cup has been held annually since 2003 in various countries around the world, providing a platform for players from different backgrounds to come together and compete in a supportive environment. The Beautiful Game highlights the importance of sports in bringing communities together and inspiring individuals to overcome adversity. The film aims to uplift and inspire viewers with its heartfelt story and message of hope.
The director and cast of The Beautiful Game aimed to pay tribute to the legacy of the Homeless World Cup and the positive impact it has had on the lives of its participants. Through authentic storytelling and performances, the film captures the challenges, relationships, and camaraderie experienced by players in the tournament. By staying true to the spirit of the Homeless World Cup, The Beautiful Game delivers a moving and inspiring cinematic experience for audiences.