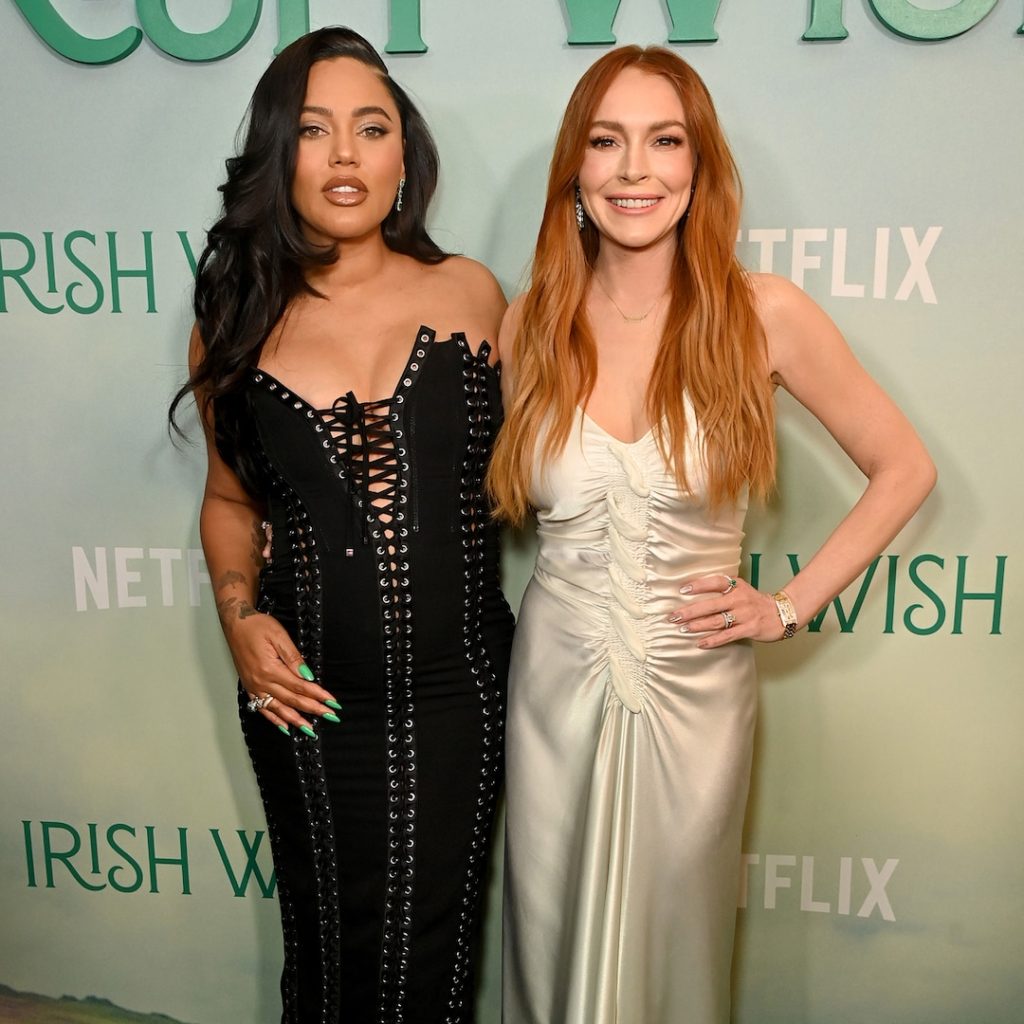
Ayesha Curry recently opened up about her long-time friendship with Lindsay Lohan, expressing admiration for the way Lohan has handled motherhood since welcoming her son Luai with husband Bader Shammas. Ayesha described Lohan as a “great mom,” noting that it has been a joy to watch her navigate this new chapter in her life. Despite being in the early days of motherhood, Ayesha praised Lohan’s ability to balance work and family life, saying that she does a great job at it.
The two women first became friends years ago and have since grown closer, with Ayesha and her husband Stephen Curry being named the godparents of baby Luai. Ayesha, who is pregnant with her fourth child, has three children of her own and knows the challenges that come with balancing family and career. She commended Lohan for being able to find a balance that works for her, even though she acknowledged that the concept of work-life balance may not truly exist.
During an exclusive interview with E! News at the Voices of Beauty Summit hosted by Landing International, Ayesha shared her admiration for Lohan’s ability to juggle motherhood and work responsibilities. She described Lohan as “killing it” in her role as a mother and applauded her for the way she has managed to find a balance between her personal and professional life. Ayesha’s words highlighted the strength of their friendship, as well as the mutual respect and admiration they have for each other.
As Lohan continues to navigate the challenges of motherhood, she has found support and love from her close friend Ayesha, who has been by her side every step of the way. The two women share a special bond that has only grown stronger over the years, and Ayesha’s role as a godparent to baby Luai demonstrates the depth of their relationship. With Lohan’s ability to find a balance between her roles as a mother and a professional, she has impressed not only her friend but also those around her, who can see the dedication and love she puts into her new role as a mother.
Lohan’s journey into motherhood has been met with admiration and respect from those around her, including Ayesha Curry, who has watched her friend navigate this new chapter with grace and strength. The bond between the two women is evident in the way Ayesha speaks about Lohan, highlighting the qualities that make her a great mother and a strong role model. With the support of her friend by her side, Lohan is sure to continue thriving in her new role as a mother, finding a balance that works for her and her family while pursuing her career goals with determination and passion.