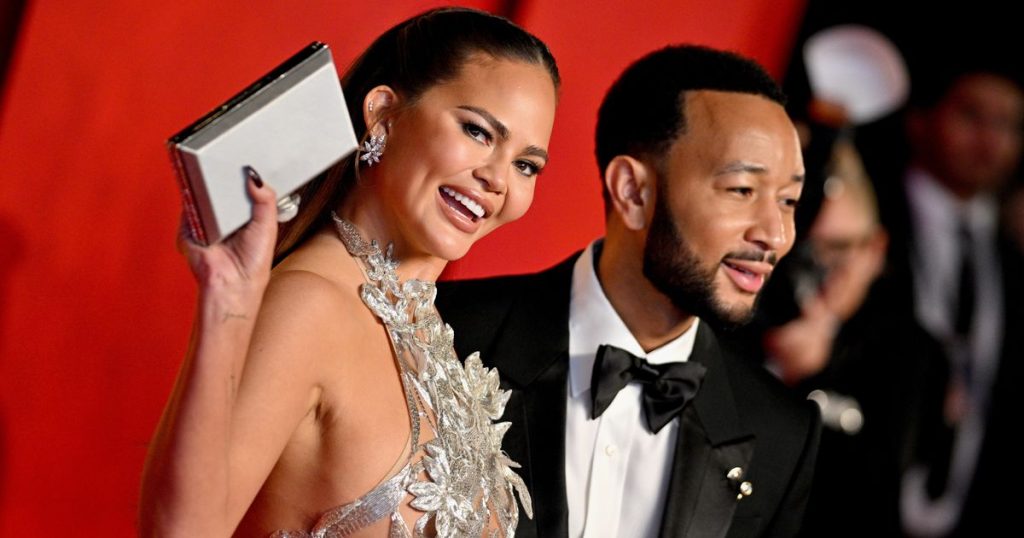
John Legend recently spoke about the importance of prioritizing his marriage and mental health with his wife, Chrissy Teigen. The couple, who have four children together, make it a point to have a staycation at a hotel once a month to take a break from their busy lives and focus on themselves and their relationship. Legend emphasized the importance of taking time for oneself and one’s relationship in order to maintain mental health and strengthen the partnership. He highlighted the need to be intentional about scheduling time for relaxation and quality time together.
Teigen, who is known for her food and lifestyle brand, Cravings by Chrissy Teigen, has been candid about the challenges the couple has faced in their relationship. In a recent interview, she shared that she struggled with jealousy and insecurity in the early years of her relationship with Legend. Teigen recounted an incident on the set of one of Legend’s music videos where she felt uncomfortable with his interaction with a female fan. However, she has since worked on overcoming these feelings and now feels more confident and secure in their relationship, viewing interactions with fans as a positive experience.
The celebrity couple’s commitment to their relationship serves as a reminder of the importance of self-care and communication in maintaining a healthy partnership. Legend emphasized the need to plan and be intentional about taking time away from the daily hustle to focus on oneself and one’s relationship. By making a monthly staycation a priority, the couple is able to recharge and reconnect, setting a positive example for others to prioritize their mental health and relationships as well. Their transparency about the challenges they have faced also helps to normalize struggles within relationships.
Teigen and Legend’s journey as a couple demonstrates that even with fame and success, relationships require effort, communication, and understanding. By sharing their experiences, they offer a relatable portrayal of the ups and downs that many couples face. Their commitment to regular check-ins and quality time together showcases the importance of investing in one’s relationship and mental well-being. Through open dialogue and a willingness to address challenges, the couple has been able to strengthen their bond and work through obstacles together.
As the couple continues to navigate the demands of their busy lives and parenting responsibilities, they serve as a reminder that self-care and prioritizing one’s relationship are essential components of a healthy partnership. By making time for themselves and each other, Teigen and Legend demonstrate the value of nurturing their connection and ensuring that their mental health remains a priority. Their commitment to growth and communication serves as a source of inspiration for others seeking to cultivate strong, resilient relationships in the face of challenges and obstacles.