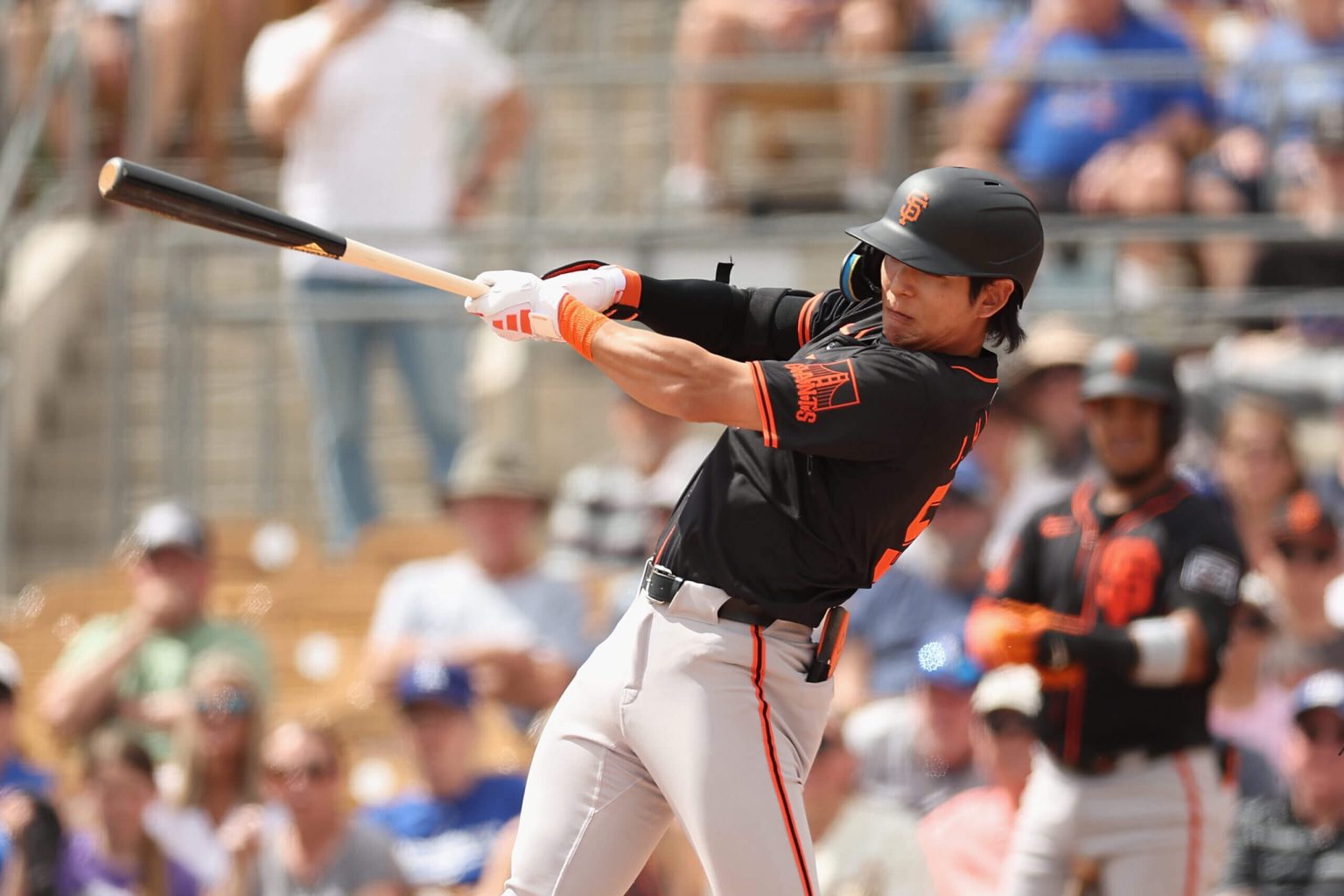
The 2024 baseball season is about to start, and a review of the previous season’s predictions for the Giants shows a mix of hits, misses, and partial credits. While some predictions were off the mark, such as Kyle Harrison making the majors and the team not taking advantage of new rules, others like Patrick Bailey’s breakout performance and the Diamondbacks winning the pennant were on point. Looking ahead to the new season, predictions include Jung Hoo Lee hitting over .300, the Giants not making significant moves at the deadline, and Luis Matos breaking the Curse of Barry Bonds in 2026.
Another prediction suggests that the Giants will continue to be cursed when it comes to having a 30-homer hitter, with Jorge Soler coming close but falling just short. Pitcher Robbie Ray is expected to struggle with his command and control upon his return, having had Tommy John surgery. Prospect Bryce Eldridge is predicted to make the top 25 of Keith Law’s top-100 list next year, showcasing his potential as a strong hitter with impressive exit velocities.
Concerns are raised about the middle-innings portion of the bullpen being an issue in the first half of the season, with a need to determine which pitchers will step up. However, the Giants are expected to have a Cy Young runner-up once again, with Logan Webb or possibly Blake Snell in the running. The prediction also suggests that the Dodgers won’t win 100 games this season, despite their strong roster and past success. Instead, the Giants are predicted to have a 92-70 record and secure a postseason spot in the 2024 season.